Materials Science
- Overview
- Members
- Announcements
- Collections
- Forum
- Projects
- Resources11
- Wiki
- Wish List
Simulation-Powered Learning Activities
Table of Contents for this page
Machine Learning for Materials Science
Kinetics
Structures
MSE educational tool: crystal structures
Educational notebooks for studying crystallography. This tool contains a few notebooks for learning crystal structures: unit cell builder, crystal builder, and a 2-D pattern builder.
MSE educational tool: visualization of stacking faults
Visualize stacking faults in FCC and BCC crystals. This notebook illustrates stacking faults in FCC (111) plane and BCC (110) plane.
MSE educational tool: X-ray diffraction (XRD) pattern
XRD pattern for BCC and FCC metals. This tool contains two notebooks: (1) Calculate and plot XRD pattern for BCC and FCC metals. (2) Calculate and plot XRD pattern for binary FCC alloys.
Visualizing Crystal Structures: An interactive group classroom activity
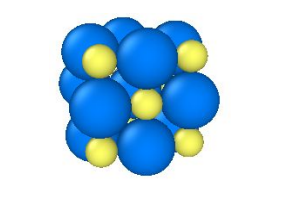
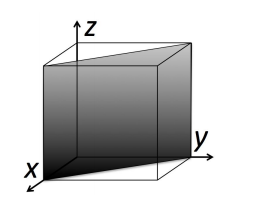
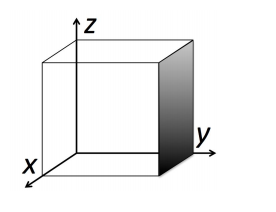
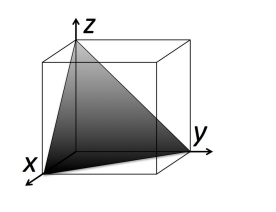

This learning activity guides students through the visualization of three-dimensional crystal structures using the software package Ovito. Students work in groups manipulating crystal structures on their personal computers, ranking the planar densities of the (100), (110), and (111) planes in face-centered cubic (FCC) and NaCl crystal structures. The activity can be performed in a 50 minute lecture session, in a lab or discussion section, or as homework.
A paper on the use of this activity in the classroom won the ASEE Materials Division Best Paper Award in 2018 (See Citation #2).
This activity can also be done using Crystal Viewer, rather than Ovito, using the following instructions:
How to Create Atomic Planes in FCC, NaCl and Simple Cubic Crystal Structures
 This step-by-step guide shows how to create atomic planes in some common crystal structures. The crystal structures can be sliced along the planes, and the atomic packing of each plane investigated.
This step-by-step guide shows how to create atomic planes in some common crystal structures. The crystal structures can be sliced along the planes, and the atomic packing of each plane investigated.
Links to saved simulations of each of the planes and structures are included.
These instructions should enable the user to create meaningful representations of many Miller planes in the numerous materials and structures available in Crystal Viewer. For example, the (111) plane in a BCC crystal is shown in the image.
Dislocation structure and propagation with molecular dynamics
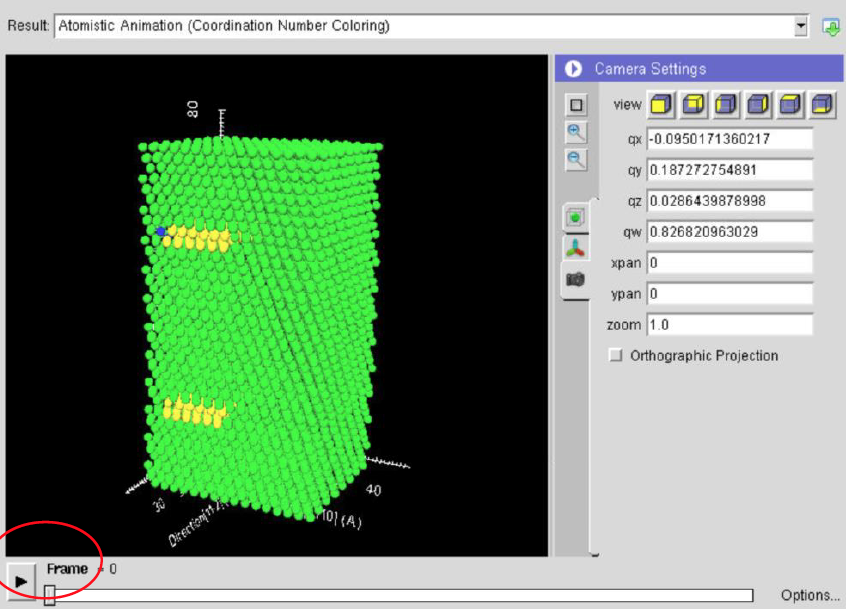 In this computational lab you will learn about dislocations via online molecular dynamics (MD) simulations using nanoHUB. The simulations involve various types of dislocations in FCC and BCC crystals. During the learning module you will be able to:
In this computational lab you will learn about dislocations via online molecular dynamics (MD) simulations using nanoHUB. The simulations involve various types of dislocations in FCC and BCC crystals. During the learning module you will be able to:
- Distinguish edge and screw dislocations in FCC and BCC crystals
- Understand the propagation of both pure dislocation types, including Burger’s vector, line direction, and slip plane
Mechanical Properties
Linear Regression Young's modulus
Use linear regression to extract Young's modulus and yield stress from stress-strain data. This tool uses python in Jupyter notebooks to analyze stress-strain data of materials and extract two materials properties: Young's modulus and yield stress. The tool has pre-loaded data, and users can also upload and analyze their own data. The tool uses Pandas to manage and organize data and scikit-learn for regression.
Users can explore, hands-on, how changing the range of strain affects the calculation of Young's moduli and script the calculations to analyze large groups of stress-strain data.
Understanding Fracture Behavior in Materials Using Cheese
 This is a resource from the Girls Learning About Materials (GLAM) outreach camp run at the University of Illinois at Urbana-Champaign. The learning objectives for this activity are:
This is a resource from the Girls Learning About Materials (GLAM) outreach camp run at the University of Illinois at Urbana-Champaign. The learning objectives for this activity are:
- Students will compare and contrast different types of fracture behavior (ductile/brittle).
- Students will understand how to characterize fracture properties.
- Students will apply their knowledge of fracture to predict how different cheeses will fail.
- Students will discuss how geometry plays a role in fracture behavior.
Nanowire Tensile Testing Laboratory
This activity compares physical tensile testing of a macroscopic sample with simulations of a tensile test on a nanowire. The simulation tool creates visualizations of the slip planes in a "slide show" of snapshots of the nanowire as the simulation progresses through elastic deformation, plastic deformation, and failure. In addition to molecular dynamics trajectories and images, stress-strain curve are generated, from which students can calculate quantities such as the modulus of elasticity, yield strength and ultimate tensile strength for the samples. This activity has been used in Purdue's undergraduate Materials Science Lab course (235) over several years, with revisions and improvements of the activity.
Three versions of this activity are available. These have employed 2 different simulation tools, the nano-Materials Simulation Toolkit and the Nanomaterial Mechanics Explorer. Both tools enable students to access advanced parameters to experiment with the simulation. Changes in the laboratory lesson were implemented after educational research was done that investigated the conceptual struggles that engineering students have with this material. Papers on educational impact of the Fall 2015 version are provided in its list of citations.
-
Original Learning Module on the Atomic Picture of Plastic Deformation in Metals
-
Uses the nano-Materials Simulation Toolkit
-
-
Fall 2014 version
-
Uses the nano-Materials Simulation Toolkit
-
-
Fall 2015 version
-
Uses the Nanomaterial Mechanics Explorer
-
Nanoscale tensile testing with molecular dynamics
 In this computational lab you will perform online molecular dynamics (MD) simulations through nanoHUB of single-crystal copper nanowires under uniaxial tension of varying orientations and analyze the results in order to:
In this computational lab you will perform online molecular dynamics (MD) simulations through nanoHUB of single-crystal copper nanowires under uniaxial tension of varying orientations and analyze the results in order to:
- Observe how slip planes in single-crystal nanowires are formed and explain plastic deformation at the atomic level in terms of dislocation motion and slip.
- Identify the characteristic features of stress-strain curves of nanoscale materials (i.e., elastic region, plastic region, Young’s modulus, yield stress)
- Differentiate plastic deformation for macro- versus nanoscale metals. Compare and contrast Young’s modulus, yield strength, and strain hardening between nanoscale, initially defect-free, single crystal samples and macroscale, polycrystalline samples
- Use critical resolved shear stress and crystallography to determine active slip systems
Ductile and brittle failure in metals with molecular dynamics
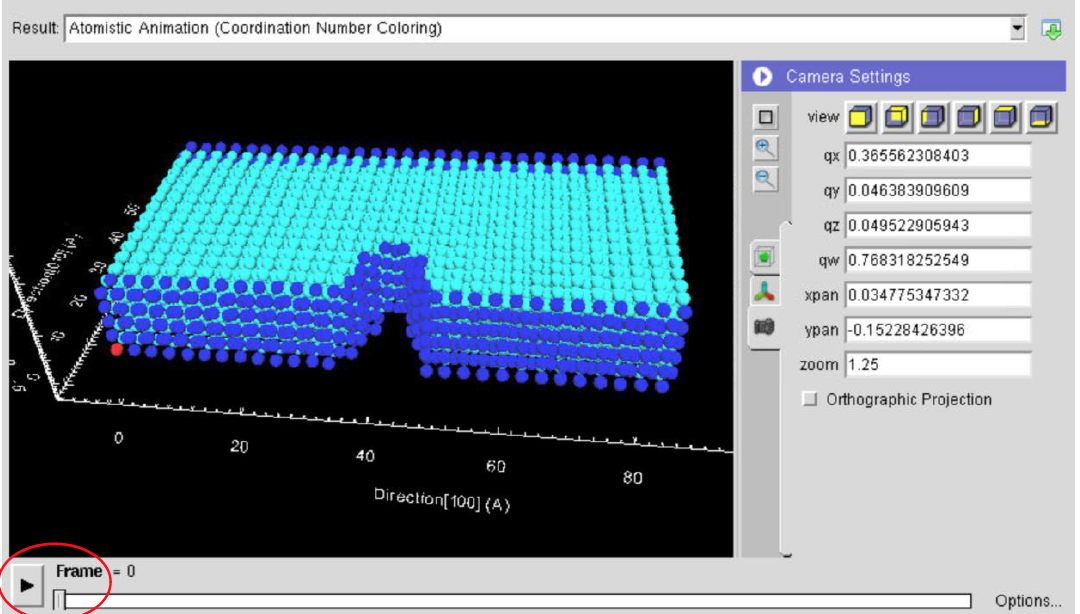 In this computational lab you will perform online molecular dynamics (MD) simulations of nanoscale cracks under uniaxial tension through nanoHUB. Simulations with varying temperature and crystal structure will provide information to:
In this computational lab you will perform online molecular dynamics (MD) simulations of nanoscale cracks under uniaxial tension through nanoHUB. Simulations with varying temperature and crystal structure will provide information to:
- Distinguish the atomistic mechanisms of ductile and brittle failure and the relationship with temperature
- Examine changes in crack propagation with crack length
MSE educational tool: elastic moduli calculations
Illustrate how to calculate elastic constants for crystalsThis notebook shows the procedure, math and 3d geometry for calculating elastic moduli.
Homework Assignment: Learning About Elastic Constants via Molecular Dynamics Simulations
In this homework assignment students will use molecular dynamics to compute the elastic constants of metals using an embedded atom model to describe atomic interactions. They will deform a single crystal along different directions and obtain c11, c12 and c44 elastic constants from the stress-strain relationships.
nanoHUB Materials Simulation Homework: Engineering the Yield Stress of a Material
This homework assignment uses the nanoplasticity lab simulation tool to enable students to explore how grain size and the competing plastic deformation mechanisms of dislocation motion and grain boundary sliding affect the yield stress of a sample. Students create conditions that lead to both the Hall-Petch and inverse Hall-Petch effects, and are asked to explain how the factors involved can lead to these two different results.
Audience: Undergraduate students learning about strengthening mechanisms and the Hall-Petch Effect.
Solving problems in Mechanics of Materials - using Jupyter Notebooks
The notebooks contain examples and tutorials to solve problems in ME 323: Mechanics of Materials.
Introduction
- Intro A brief Python tutorial
- Linear Equations Intro (Octave)
- An introduction to solving systems of linear equations (Octave)
Examples
- Indeterminate Rod (Octave) Solution of Indeterminate Rod Problem (Octave)
- Shear and Bending Diagrams (Octave)
- Beam Shear & Bending Diagram Tool (Octave)
- Mohr's Circle (Octave) Mohr's Circle Notebook (Octave)
- Mohr's Circle (Python) Mohr's Circle Notebook (Python)
- Interactive Mohr's Circle Notebook containing an interactive Mohr's Circle (Python)
Finite Element Method
- 1D Rod Introduction Introduction to the following 1D FEM Scripts
- 1D Rod (Octave) 1D Finite Element Script (Octave)
- 1D Rod (Python) 1D Finite Element Script (Python)
This tutorial is intended for use in understanding and practicing the basics of the finite element method (FEM) in one dimension (1D). The scripts in these notebooks solve the one-dimensional problem described below. The notebooks allow these scripts to be modified, run, and downloaded.
One of the notebooks is written in Octave, which is similar to Matlab. The remaining script is written in Python 3.0. The scripts are functionally identical otherwise.
The scripts solve for the displacement u of the bar loaded as shown in Figure 1. Both the analytical and numerical solutions are presented in this resource.

Figure 1: A bar of cross-sectional area A, Young's modulus E, and length L, fixed at the left end and loaded with a force F at the right end. The displacement u at the right-hand side is to be determined.

Figure 2: Division of the rod into N elements and N+1 nodes, with each element represented by an equivalent spring of stiffness ki.
Machine Learning for Materials Science
See the Machine Learning Group for more resources
Tools, tutorials and seminars for Machine Learning.
Machine Learning for Materials Science, Part 1
Data science and machine learning are playing an increasingly important role in materials science and engineering. This tutorial provides examples of the use of these tools in the field of materials science using Jupyter notebooks that contain step-by-step explanations of each activity using live code that can be easily run, modified and tested to aid in learning.
The initial set of tutorials focus on:
i) Data query, organization and visualization,
ii) Developing a simple model using linear regression to explore correlations between materials properties, and
iii) Neural network (NN) models trained to predict materials properties, such as the modulus of elasticity, from basic element properties.
iv) NN models to classify materials; predicting their crystal structure based on other elemental properties.
Machine Learning Lab Module
This lab activity gives an overview of a materials science machine learning workflow. It starts with initial data cleaning and ends with analyzing various predictions the model could make. Supplementary material is included so that this can be used as a one week module for introducing students to machine learning in materials science.
Tensor Flow Tutorial
This is a set of tutorials using Jupyter notebooks in nanoHUB that will get you started with machine learning using TensorFlow and Keras. The set of tutorials was taken from TensorFlow (https://www.tensorflow.org/tutorials/) with copyright by François Chollet and is deployed with minimal modifications. Using nanoHUB resources users can run the tutorials, modify them and explore machine learning from any laptop or tablet, without downloading or installing any software.
Kinetics
Computational Labs in Kinetics of Materials and Process Design (California Polytechnic State University)
This resource contains three practical labs that use various nanoHUB simulation tools for (a) Diffusion in Solids, (b) Nucleation, Crystallization, and Growth, and (c) Phase Transformation.
Exploration of the Oxidation Rate of Silicon Wafers via Simulation
This teaching resource is designed for instructors who would like to introduce exploration through simulation into their lessons on silicon oxidation. Step-by-step instructions are provided for running the process lab oxidation simulation, and guiding questions are provided that should help students discover important trends in the oxidation rate when different process parameters are changed.
Thermodynamics and Phase Transitions
Melting via molecular dynamics simulations
In this assignment you will use MD simulations to study melting in metals using the nanoMATERIALS simulation tool in nanoHUB. You will create atomistic models for bulk Al and a nanoparticle and use MD simulations to heat them up and study their evolution. You can visualize the atomic configuration as the temperature is increased and after melting. From the results of the simulation you can obtain the melting temperature and the heat of fusion.
Melting with molecular dynamics
In this computational lab you will perform online molecular dynamics (MD) simulations using the nanomaterial mechanics explorer simulation tool to melt nickel samples and analyze the results in order to:
- Understand the process of melting at atomic scales
- Identify effects of surfaces and specimen size
- Describe differences between simulation and experiment
Diffusion and Transport Properties
2D Diffusion Game
The Diffusion Game package introduces students to computational materials modeling by simulating the 2D diffusion of metal atoms. See the Supporting Docs tab for more information, and a paper game board.
Electronic Structure and Properties
Computer Modeling Module: Chemical Reaction Simulation using SIESTA
This activity guides students through a module using the SIESTA DFT tool that is housed within the MIT Atomic Scale Modeling Toolkit on nanoHUB. Instructional videos, background reading, reminders and the assignment are included.
Online Simulation tutorial and assignment: bonding curves in H2 and He2 molecules
In this tutorial students will use density functional theory (DFT) calculations using the nanoHUB tool SeqQuest to study bonding in two simple molecules: H2 and He2. The tutorial shows how to compute energy as a function of bond distance and extract the equilibrium bond distance and bond strength.
Disclaimer: While very powerful, DFT makes well-known approximations and the results obtained in this module are approximate.
Online Simulation tutorial and assignment: electronic structure and spin of the O atom
In this tutorial students will use density functional theory (DFT) calculations using the nanoHUB tool SeqQuest to study the electronic structure of the oxygen atom. The tutorial shows how to compute energy for the spin 1 (triplet) and spin 0 (singlet) states and analyze the exchange energy.
Disclaimer: While very powerful, DFT makes well-known approximations and the results obtained in this module are approximate.
Online Simulation tutorial and assignment: electronic structure and spin in O2 molecule
In this tutorial students will use density functional theory (DFT) calculations using the nanoHUB tool SeqQuest to study the electronic structure of the oxygen atom. The tutorial shows how to compute energy for the spin 1 (triplet) and spin 0 (singlet) states and analyze the exchange energy.
Disclaimer: While very powerful, DFT makes well-known approximations and the results obtained in this module are approximate.
Using DFT to Predict the Equilibrium Lattice Parameter and Bulk Modulus of Crystalline Materials
This activity guides users through the use of DFT calculations with Quantum ESPRESSO in nanoHUB to calculate the total energy of a crystal structure. By varying the volume of the structure, and calculating the associated energies, the equilibirum structure can be found.
Users are guided through the use of the Materials Project to obtain structural information to enter into the Quantum ESPRESSO simulation tool.
Using DFT to Simulate the Band Structure and Density of States of Crystalline Materials
In this activity, DFT is used to simulate the band structure and density of states of several crystalline semiconductors. Users are instructed in how to use the Bilbao Crystallographic Server to select a path through the Brillouin zone for each structure.
Bonding and Bandstructure in Silicon
This Learning Module contains Introductory Lectures, Tutorials and an Online-simulation Lab Assignment that allows students to explore how electronic bands form in silicon and other materials. Students will perform online density functional theory calculations and explore the band structure and bonding of various materials.
Audience: undergraduate and graduate students as well as instructors interested in electronic properties of materials.
Learning Module: Band Structure for Pure and Doped Silicon
In this lab, students will learn to perform online density functional theory (DFT) simulations to compute band structures and density of states (DOS) for pure and doped Si using the DFT Material Properties Simulator available on nanoHUB.
The students will work with crystalline pure and doped Si. They will analyze the results to:
- Determine the nature of the system (metallic, semiconducting or insulating)
- Interpret the DFT predicted band structure and density of states (DOS)
- Understand variation in band structure and DOS for doped semiconductor
